Single-cell transcriptomics reveal how root tissues adapt to soil stress
Mingyuan Zhu, Che-Wei Hsu, Lucas L Peralta Ogorek, Isaiah W Taylor, Salvatore La Cavera, Dyoni M Oliveira, Lokesh Verma, Poonam Mehra, Medhavinee Mijar, Ari Sadanandom, Fernando Perez-Cota, Wout Boerjan, Trevor M Nolan, Malcolm J Bennett, Philip N Benfey, Bipin K Pandey
Nature; 2025 Jun; 642(8068):721-729. doi: 10.1038/s41586-025-08941-z.
Abstract
Land plants thrive in soils showing vastly different properties and environmental stresses1. Root systems can adapt to contrasting soil conditions and stresses, yet how their responses are programmed at the individual cell scale remains unclear. Using single-cell RNA sequencing and spatial transcriptomic approaches, we showed major expression changes in outer root cell types when comparing the single-cell transcriptomes of rice roots grown in gel versus soil conditions. These tissue-specific transcriptional responses are related to nutrient homeostasis, cell wall integrity and defence in response to heterogeneous soil versus homogeneous gel growth conditions. We also demonstrate how the model soil stress, termed compaction, triggers expression changes in cell wall remodelling and barrier formation in outer and inner root tissues, regulated by abscisic acid released from phloem cells. Our study reveals how root tissues communicate and adapt to contrasting soil conditions at single-cell resolution.
See https://pubmed.ncbi.nlm.nih.gov/40307555/
![]()
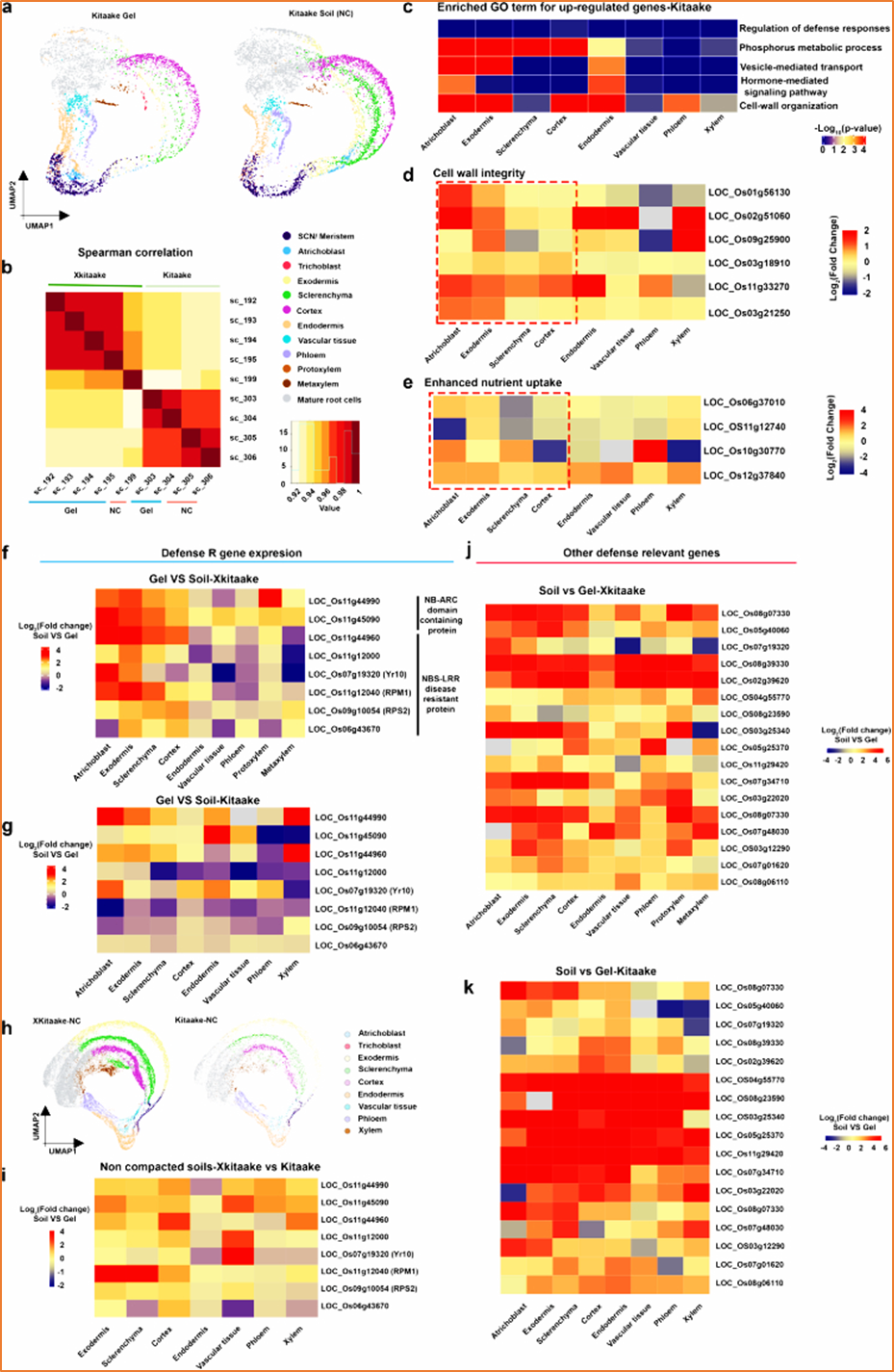
Fig. 4. The XA21 transgene in the Xkitaake background does not alter overall gene expression patterns when root growth conditions change from gel to soil conditions.
Views: 240


