Integrating image-based phenotyping and QTL mapping to enhance genetic resistance and accelerate breeding for bacterial grain rot resistance in rice
Jae-Ryoung Park, Gileung Lee, Hyun-Sook Lee, Seung Young Lee, Su-Kyung Ha, Kyeongmin Kang, Mina Jin, Jung-Pil Suh, Jeonghwan Seo, Young Mi Choi, Bong Choon Lee & Hyun-Su Park
TAG; March 26 2026; vol. 139; article 108
![]()

Figure: Rice bacterial grain rot disease by Burkholderia glumae
Abstract
Bacterial grain rot (BGR), caused by Burkholderia glumae, is a major disease that reduces the yield of rice (Oryza sativa L.), thereby threatening food security. Conventional phenotypic analysis methods face limitations in objectively evaluating disease resistance and understanding the genetic basis. In this study, we integrated image-based phenotypic analysis with QTL mapping to screen for QTLs and candidate genes associated with B. glumae resistance. B. glumae was inoculated into 189 recombinant inbred lines (RILs) derived from Kele (resistant) and IS592BB (susceptible), followed by visualization and quantitative analysis using DAB staining. Phenotypic parameters, including the field resistance score, ratio of diseased spikelets (%), DAB staining intensity, and ratio of diseased area (%), were measured and used for QTL mapping. On chromosome 1, within Chr01_24592710-Chr01_37274755, four QTLs—qFRS1 [LOD: 5.98, phenotype variation explained (PVE): 15.41%], qRDS1 (LOD: 5.29, PVE: 18.56%), qQDS1 (LOD: 9.58, PVE: 22.02%), and qRDA1 (LOD: 8.44, PVE: 31.51%)—were identified as overlapping. After fine-mapping we narrow down Chr01_33472174-Chr01_33838140 and a total of 16 candidate genes were screened this region. Among which OsBGq1 was found to encode a nucleotide-binding LRR receptor (NLR) domain. OsBGq1 expression increased significantly upon B. glumae infection. Additionally, RILs Kele type of Chr01_33472174-Chr01_33838140 presented increased ROS-scavenging enzyme activity and phytoalexin accumulation upon B. glumae infection, contributing to increased resistance. The integration of DAB-based quantitative phenotyping with QTL mapping is proposed to provide a more objective indicator for identifying genes associated with resistance to BGR.
See: https://link.springer.com/article/10.1007/s00122-026-05216-7
![]() Figure 2
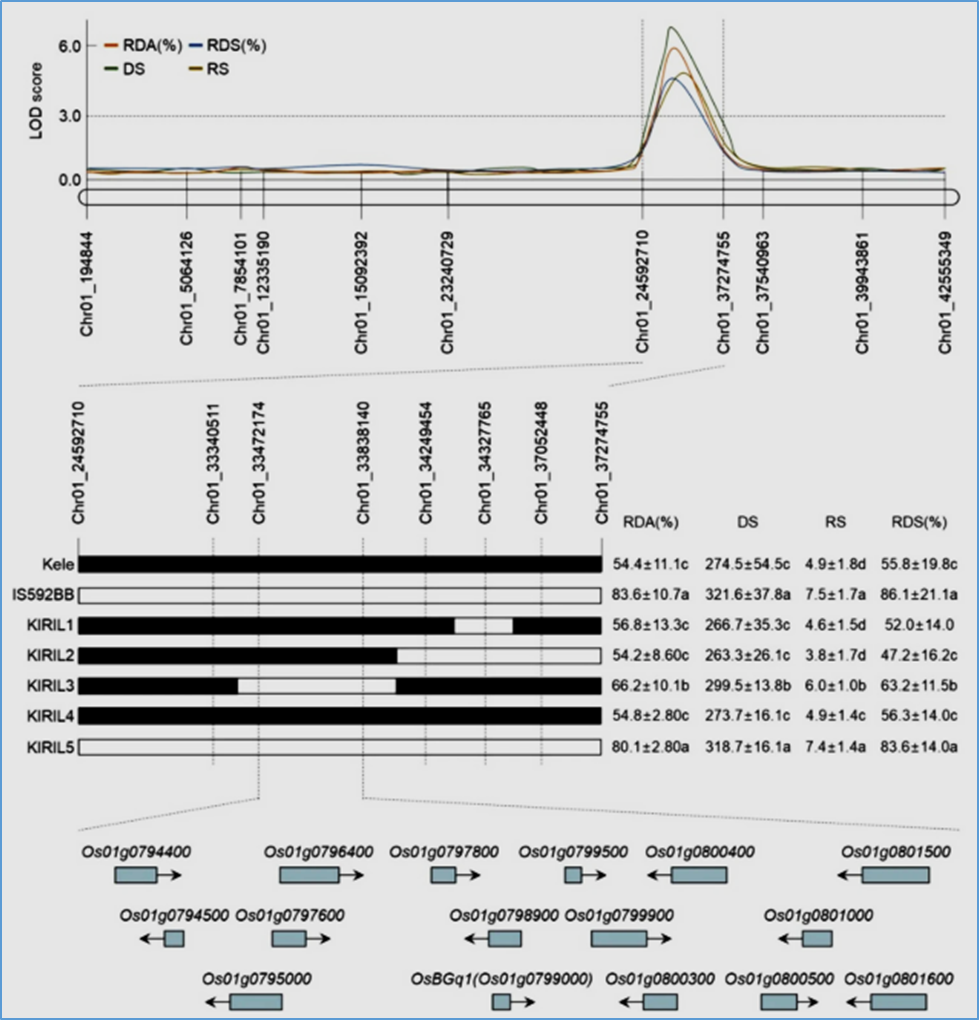
Figure 2
Fine mapping of the Chr01_24592710-Chr01_37274755 region on chromosome 1 in KIRILs to screen for candidate genes associated with B. glumae resistance. The black bars represent chromosome segments derived from Kele (resistant rice cultivar), whereas the white bars indicate segments inherited from IS592BB (susceptible rice cultivar). Six additional markers (Chr01_33340511, Chr01_33472174, Chr01_33838140, Chr01_34249454, Chr01_34327765, and Chr01_37052448) were incorporated within the Chr01_24592710– Chr01_37274755 region. In KIRILs, these markers identified five distinct genotypes. KIRIL1, KIRIL2, KIRIL3, KIRIL4, and KIRIL5 are representative recombinant inbred lines derived from the Kele and IS592BB cross, each carrying a distinct recombined genotype within the KIRIL population. Following B. glumae infection, phenotype parameters (RDA, DS, RS, and RDS) were measured and are summarized to the right of the genotype information. This figure illustrates the distribution of 16 candidate genes located within the Chr01_33472174– Chr01_33838140 region. The size of the blue boxes represents the relative length of the genomic DNA sequence, whereas the direction of the arrows indicates the transcriptional orientation of the candidate genes. The data are shown as the means ± SDs from five independent biological replicates per line. Means sharing the same letter are not significantly different (P < 0.05), as determined by Duncan’s multiple range test
Views: 35


