Deciphering cassava brown streak virus infection in cassava through VPg mediated host protein interactions
Sumesh M Kakkunnath, Sophie Bouvaine, Siji P Kavil, M N Maruthi
Arch Virol.; 2026 Mar 19; 171(4):135. doi: 10.1007/s00705-026-06571-y.

![]()
Abstract
Cassava brown streak disease (CBSD) poses a major threat to cassava production in Africa. Identifying cassava proteins that interact with cassava brown streak virus (CBSV), the major causal virus, can help elucidate the mechanisms of infection and resistance. Here we constructed a cassava cDNA library and screened for proteins interacting with CBSV viral genome-linked protein (VPg) using yeast two-hybrid assays, identifying 36 interactors. Four candidates were validated in Nicotiana benthamiana via bimolecular fluorescence complementation. Functional categories included chloroplast proteins, ribosomal components, chaperones, metabolic enzymes, and defence-related proteins. In prior RNA-seq datasets from CBSV or UCBSV-inoculated cassava, comprising the susceptible variety Albert and the resistant variety Namikonga, 16 VPg-interacting genes were identified among the differentially expressed genes (|log₂FC| > 1), with 2 detected in Albert and 15 in Namikonga. These results indicate that VPg-interacting host proteins are involved in CBSV infection dynamics and also explaining the differential responses of resistant and susceptible cassava varieties, thus offering potential new molecular targets for CBSD management.
See: https://pubmed.ncbi.nlm.nih.gov/41851384/
![]()
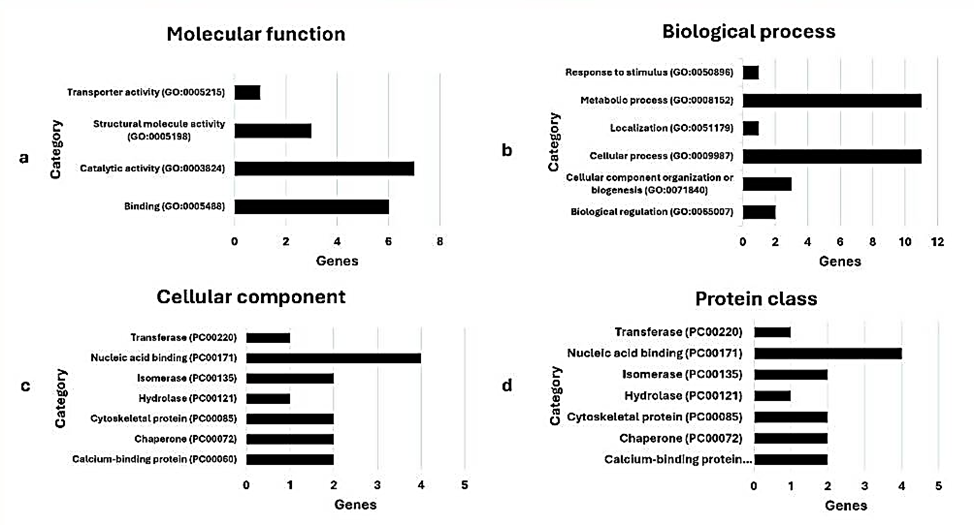
Figure 2: Gene ontology based on (a) molecular function; (b) biological process; (c) cellular component; and (d) protein class
Views: 5


