Genomic approaches to build de novo elite breeding gene pools from locally adapted landraces
Safiétou Tooli Fall, Alexander Kena, Brian R. Rice, Ghislain Kanfany, Cyril Diatta, Ndjido A. Kane, Allan K. Fritz & Geoffrey P. Morris
Theoretical and Applied Genetics; January 7 2026; vol. 139; article 28
![]() Abstract
Abstract
Many nascent breeding programs aim to achieve genetic gain by crossing locally-elite germplasm, but a lack of systematic approaches to develop elite gene pools from locally adapted varieties hinders their progress. Motivated by the observation of undesirable transgressive segregation in presumed elite crosses in Senegalese cereal breeding programs, we designed approaches for de novo development of elite gene pools from locally adapted landrace-derived germplasm. We first define two types of “elite” germplasm: iso-elite, phenotypically similar and genetically homogeneous for locally adapted traits (“attained traits”); versus allo-elite, phenotypically similar, but genetically heterogeneous for attained traits. Next, we defined two genomic approaches for de novo inference of elite gene pools: population-based genotypic inference (PGI) and QTL-based genotypic inference (QGI), and compared to a family-based phenotypic inference (FPI) approach. Using simulations that trace the evolution from locally adapted landraces to elite breeding lines, we evaluate the effectiveness of these strategies in nascent forward breeding programs. QGI accurately and cost-effectively identifies both iso- and allo-elite pairs, regardless of the underlying trait architecture, while PGI is less sensitive when trait architecture is oligogenic. Over ten cycles of phenotypic recurrent selection, programs based on iso-elite crosses consistently outperformed those based on allo-elite crosses for genetic gain. The findings highlight the value of trait genetic architecture knowledge for elite gene pool development and provide a practical roadmap for elite germplasm development in modernizing breeding programs.
See https://link.springer.com/article/10.1007/s00122-025-05124-2
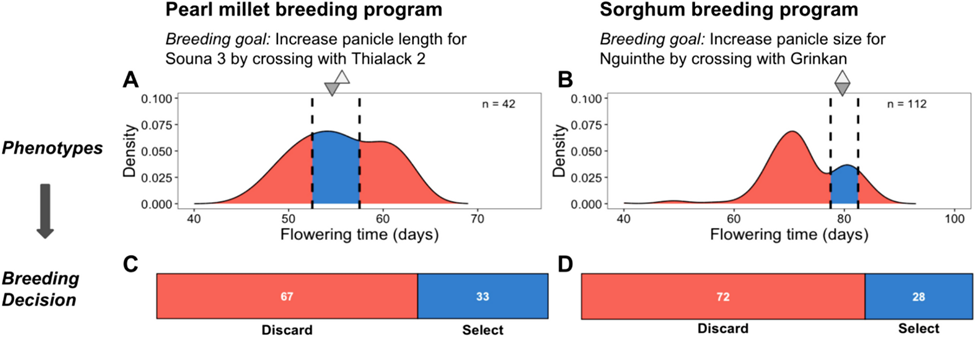
![]() Figure 1: From Genomic approaches to build de novo elite breeding gene pools from locally adapted landraces.
Figure 1: From Genomic approaches to build de novo elite breeding gene pools from locally adapted landraces.
Undesirable transgressive segregation for an attained trait in progeny from presumed elite-by-elite crosses substantially reduces the number of candidates for selection. Phenotypic distribution of flowering time in progeny from presumed elite-by-elite crosses in millet (A; F5 progeny) and sorghum (B; F7 progeny) breeding programs in Senegal. For pearl millet, Souna 3 was crossed with Thialack 2 to improve panicle length while maintaining earliness and high yield. For sorghum, Nguinthe was crossed with Grinkan to improve panicle size while maintaining lateness. Flowering time is an attained trait in these programs, since the parents (triangles) are within the acceptable range of phenotypes (black dotted lines) according to the breeding product profiles. For both crops, substantial transgressive segregation was observed, with few progeny within the acceptable range (blue shading in A and B), and most progeny falling outside (red shading in A and B). Accordingly, the percentage progeny of discarded (red in C and D) versus selected (blue in C and D) progeny was high, with only 33 and 28% of the progeny for pearl millet and sorghum, respectively, selected as acceptable with respect to flowering time
Views: 53


