Fine mapping and transcriptomics reveal OSG function in regulation of grain size and pollen fertility in rice (Oryza sativa)
Sijie Zhao, Siyue Zhang, Weichen Xu, Sijia Fang, Mei Liu, Lei Wang, Xianyang Zhang, Bingxu Chen, Shuya Wei, Heming Zhao
PLoS One; 2026 Jan 8; 21(1):e0338401. doi: 10.1371/journal.pone.0338401.
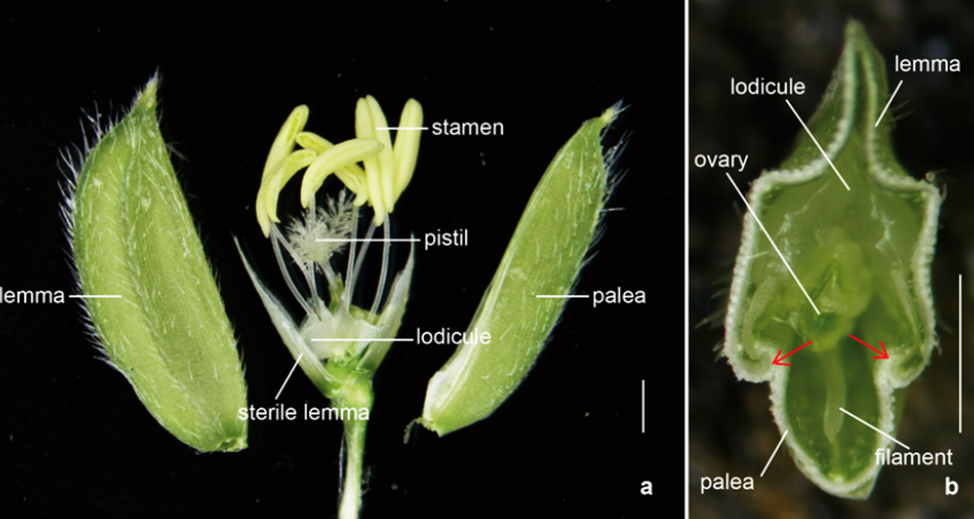

Figure: The spikelet structure in rice (Wang et al. 2023).
Abstract
Grain shape is a critical factor that directly influences rice yield and quality, however, the molecular mechanisms underlying the regulation of grain shape development remains elusive. In this study, we characterized an oval-shaped grain mutant, osg, from the rice radiation mutagenesis mutant library via 60Co-γ ray irradiation. Compared to ZH11, the osg mutant exhibited decreased grain length and thousand-grain weight but increased grain width and thickness, while displaying significantly reduced plant height, tiller number, pollen viability, and seed setting rate. Map-based cloning revealed that OsSRS3, a known regulator of rice grain size and fertility, was identified as the candidate gene for OSG. The loss-of-function mutant in OsSRS3 exhibit abnormal phenotypes similar to that in osg. Comparative transcriptome sequencing of young panicles from ZH11 and osg showed that up-regulated genes were predominantly enriched in pathways related to plant hormone signal transduction and MAPK signaling, whereas down-regulated genes were mostly associated with starch and sucrose metabolism. Further analysis of differentially expressed genes (DEGs) within the pathways revealed that multiple DEGs in osg, such as OsJAZ11, OsUgp1 and OsUgp2, have been functionally characterized, and their mutant phenotypes align with those observed in osg. These findings provide a crucial foundation for elucidating the molecular mechanisms by which the SRS3 gene regulates grain size and fertility in rice.
See: https://pubmed.ncbi.nlm.nih.gov/41505466/
![]()
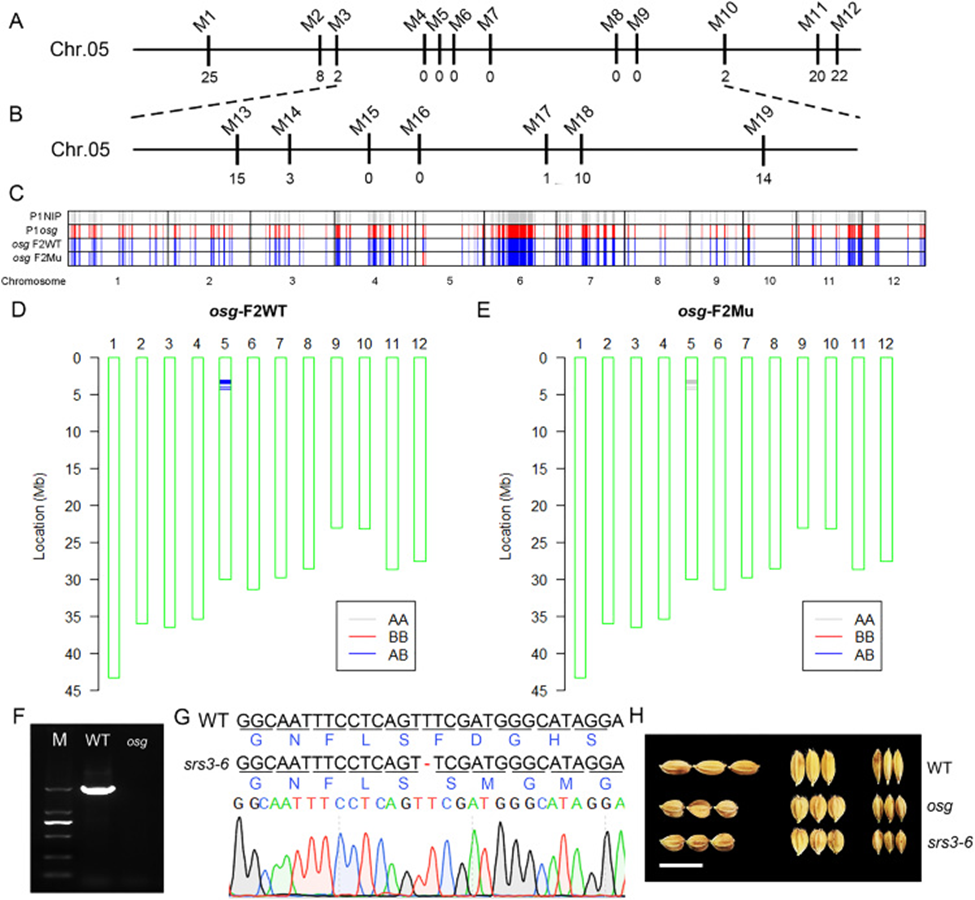
Figure 5: Localization of the OSG gene and analysis of candidate genes.
(A) Preliminary localization of the OSG gene using InDel markers M1 and M12 on chromosome 5; (B) Fine mapping of the OSG gene using newly designed InDel markers between M3 and M10, further narrowed down to between M14 and M17; (C) Genotype analysis of parents and F2 pools using SNP chip data; parent osg appears gray (homozygous), parent NIP appears red (homozygous), indicating homozygous genotypes for both parents; (D) SNP analysis at differential loci showing mostly heterozygous genotypes in the WT pool; (E) SNP analysis at differential loci showing homozygous mutant genotype (AA) in the mutant pool; (F) Amplification abnormalities observed in exons 8, 9, 10 and 11 of the SRS3 gene in osg; (G) Identification of a homozygous individual plant srs3-6 with a single-base deletion (T) leading to loss of function in the SRS3 gene; (H) Observation of grain morphology in srs3-6, scale bar = 1 cm.
Views: 57


