A natural variant of an MYC2 gene in soybean contributes to resistance against the common cutworm
Xiao Li, Dezhou Hu, Zhongyi Yang, Linyan Cai, Mengshan Zhang, Hailun Liu, Dongquan Guo, Shupeng Dong, Changyun Yang, Fang Huang, Deyue Yu, and Hui Wang
PNAS; December 26 2025; 122 (52) e2424956122; https://doi.org/10.1073/pnas.2424956122
Significance
Plants are constantly threatened by herbivores during their life cycle. In soybean, the common cutworm (CCW) is a major pest responsible for significant yield losses in many regions. For example, in Asia, its occurrence frequently reduces annual yields by 10 to 20%, with losses exceeding 50% during severe outbreaks. The lack of sufficiently effective defense genes limits soybean breeding for resistance to this insect. This study describes the cloning and characterization of a key defense gene, GmMYC3, in soybean that confers resistance to the CCW. Omics analyses revealed that GmMYC3 affects multiple defense-related genes and metabolites, such as soybean trypsin inhibitors. These findings identify GmMYC3 as a promising genetic resource for breeding soybean plants with enhanced resistance to the CCW.
Abstract
Herbivory is destructive for crop production in many regions worldwide. The induced plant response to herbivores promotes resistance; therefore, characterizing the mechanisms underlying natural host resistance is highly important. However, the genetic components of resistance to herbivores in the staple food crop soybean remain elusive. Here, a key defense gene, GmMYC3, was identified via joint linkage and association mapping in soybean. GmMYC3 encodes an MYC2 transcription factor that is rapidly activated after jasmonic acid treatment or herbivore attack and confers resistance to a major pest, the common cutworm (CCW), in soybean. GmMYC3 positively regulates multiple biotic stress-related genes, among which GmMYC3 triggers high expression of Kunitz-type trypsin inhibitors alongside its homolog GmMYC1 and downstream GmWRKY56. GmMYC3 overexpression results in massive accumulation of trypsin inhibitors in soybean leaves, interferes with the protein digestion and absorption function in larvae that are fed these leaves, and retards larval and pupal development of the CCW. The results of the field tests of the transgenic plants corroborate the defense role of GmMYC3. Evolutionary and population genetic analyses suggest that the elite haplotype of GmMYC3 contributes to resistance against the CCW without significant reduction in seed yield and quality. Notably, this haplotype appears at a low frequency in domesticated germplasms. This study sheds light on the molecular mechanism underlying plant resistance to the CCW and provides potentially valuable resources for breeding soybean plants with elevated resistance against this pest.
See https://www.pnas.org/doi/10.1073/pnas.2424956122
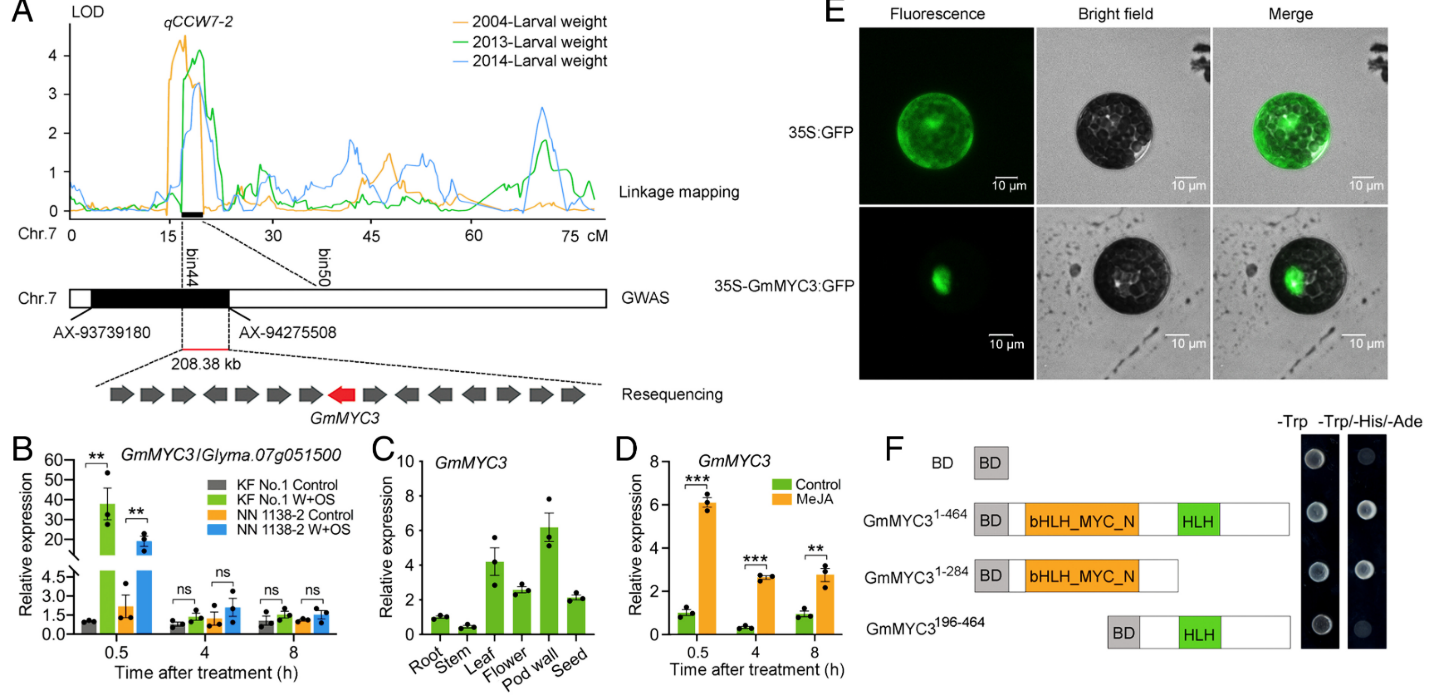
FIGURE 1: Identification of GmMYC3 as a candidate gene that regulates soybean resistance to the CCW. (A) GmMYC3 was identified by combining linkage mapping, GWAS, and resequencing analysis. The arrows represent genes. The direction of the arrow represents the forward and reverse genes. Chr, chromosome; LOD, logarithm of odds. (B) Relative expression of GmMYC3 in the parental lines KF No. 1 (resistant) and NN 1138-2 (susceptible) after W+OS treatment (n = 3). Leaves were treated without and with W+OS, and after 0.5, 4, and 8 h, samples from both the control and treatment plants were collected for qRT–PCR analysis. (C) Relative expression of GmMYC3 in different tissues of KF No. 1 (n = 3). (D) Relative expression of GmMYC3 in the leaves of KF No. 1 at 0.5, 4, and 8 h after MeJA treatment compared with the control conditions (n = 3). For (B–D), the value of each plant is represented by a point. A two-tailed t test was used for statistical analyses: **P < 0.01; ***P < 0.001; ns, not significant. The error bars denote ±SEM. (E) Subcellular localization of the GmMYC3-GFP fusion protein in Arabidopsis protoplasts. GFP alone was used as the control. (Scale bar, 10 μm.) (F) Transcription activity of full-length, N-terminal, or C-terminal GmMYC3 in yeast. −Trp, synthetic dropout medium lacking tryptophan; −Trp/−His/−Ade, synthetic dropout medium lacking tryptophan, histidine, and adenine. The superscript numbers indicate the GmMYC3 amino acid residues used for vector construction. GAL4 DNA BD alone was used as the negative control.
Views: 177


