A Non-Host Pathogen Elicitor Induces Blast Resistance Mediated by OsNAC78-Pir7b Module in Rice
Yunjie Xie, Yuling Lai, Xingqian Wu, Xiangzhen Yu, Jingwen Wei, Mengge Chen, Yongsheng Zhu, Liping Chen, Hongguang Xie, Qiuhua Cai, Huaan Xie, Jianfu Zhang
Plant Cell Environ.; 2026 Mar 29. doi: 10.1111/pce.70500. Online ahead of print.
![]()
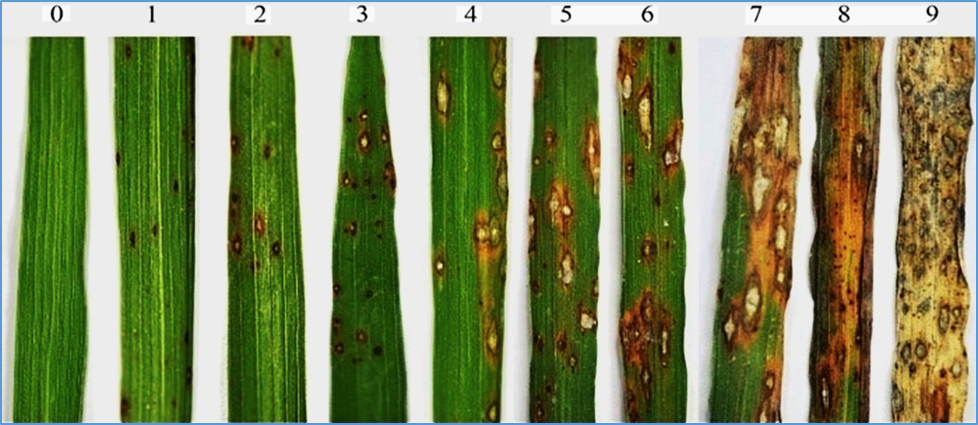

Abstract
Plants exhibit broad-spectrum and persistent resistance induced by non-host pathogens. Previous studies have found that syringolin A secreted by Pseudomonas syringae pv. syringae can activate the expression of defense-related gene Pir7b in non-host rice, but the underlying mechanism remains ambiguous. In this study, we found that OsNAC78, a transcription factor upstream of Pir7b, is possible to participate in the immune pathway. First, the OsNAC78 transgenic plants were inoculated with Magnaporthe oryzae, showing that OsNAC78 is a positive regulator for blast resistance. Meanwhile, the transgenic lines overexpressing or knocking out Pir7b were constructed and evaluated for resistance, displaying that Pir7b confers blast resistance. Subsequently, OsNAC78-Pir7b module is activated by enhancing activity of OsNAC78 and Pir7b promoters under the syringolin A treatment. The transcriptional activation of Pir7b is blocked in the absence of OsNAC78, indicating that OsNAC78 is a key regulator of Pir7b expression. Further, it was found that syringolin A can activate the strongest resistance in the wild type, while the weakest in the Osnac78/pir7b mutant, suggesting that syringolin A induces rice immunity through the OsNAC78-Pir7b module. Overall, we discovered that syringolin A secreted by non-host pathogens can induce the expression of the transcription factor OsNAC78 in rice, subsequently upregulate the transcription of the downstream gene Pir7b, and ultimately trigger blast resistance by ROS accumulation. This study reveals how the secretion of non-host pathogen stimulates plant immune system, providing new insights for plant disease control.
See https://pubmed.ncbi.nlm.nih.gov/41906318/
![]()
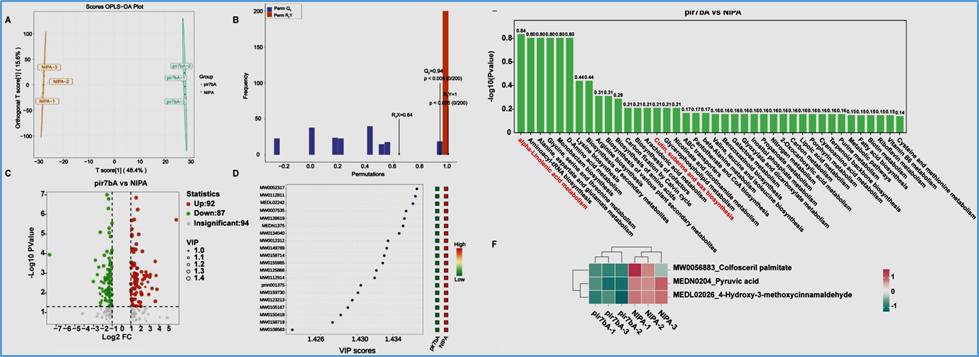
Figure 8
Non-targeted metabolomics analysis of pir7bA and NIPA. (A) The OPLS-DA score plot for pir7bA_vs_NIPA. The horizontal axis direction indicated differences between groups; the vertical axis represents orthogonal principal components, with its direction revealing differences within groups. (B) OPLS-DA validation plot. R2X and R2Y denote the explanatory power of the constructed model for the X and Y matrices, respectively, while Q2 indicates the model's predictive capability. The closer these three metrics are to 1, the more stable and reliable the model is. A Q2 value greater than 0.5 indicates an effective model, while a Q2 value greater than 0.9 signifies an excellent model. (C) Volcano plot of differentially expressed metabolites between pir7bA and NIPA. Green dots represent downregulated metabolites, red dots represent upregulated metabolites, and gray dots indicate metabolites detected but not significantly different. (D) VIP Value Plot + Heatmap (VIP > 1) displayed only the top 20 differentially expressed metabolites. (E) Bar chart showing KEGG pathway enrichment significance for differential metabolites, displaying the top 40 pathways. Pathways annotated in red were those associated with ROS activity. (F) Heatmap of the contents of MW0056883_Colfosceril palmitate, MEDN0204_Pyruvic acid, and MEDL02026_4-Hydroxy-3-methoxycinnamaldehyde. [Color figure can be viewed at wileyonlinelibrary.com]
Views: 10


